Quick Tutorial¶
Installation/Usage¶
As the package has not been published on PyPi yet, it CANNOT be install using pip.
For now, the suggested method is to put the file Complex_Contagions.py in the same directory as your source files and call from Complex_Contagions import geometric_network
Initiate a geometric_network object¶
Create a geometric network on a ring. Band_length corresponds to the number of neighbors to connect from both right and left making the geometric degree 2*band_length
n = 20
d2 = 2
ring_latt= geometric_network('ring_lattice', size = n, banded = True, band_length = 3)
Add noise to geometric network¶
Use add_noise_to_geometric() method to manipulate the network topology. The second parameter describes the non-geometric degree of every node.
ring_latt_k_regular.add_noise_to_geometric('k_regular', d2)
Sample Excitation Simulation¶
Run the complex contagion on the network we have created. Key parameters are threshold, C and $alpha = frac{nGD}{GD}$
n = 200
d2 = 2
ring_latt_k_regular = geometric_network('ring_lattice', size = n, banded = True, band_length = 3)
ring_latt_k_regular.add_noise_to_geometric('k_regular', d2)
T = 100 # number of iterations
seed = int(n/2) # node that the spread starts
C = 1000 # Geometrically, this describes the turning of the sigmoid function
threshold = 0.3 # resistence of the node to it's neighbors' excitation level
Trials = 2 # number of trials
refractory_period = False ## if a neuron is activated once, it stays activated throughout.
fig, ax = plt.subplots(Trials,1, figsize = (50,10))
first_excitation_times, contagion_size = ring_latt_k_regular.run_excitation(Trials, T, C, seed, threshold, refractory_period, ax = ax)
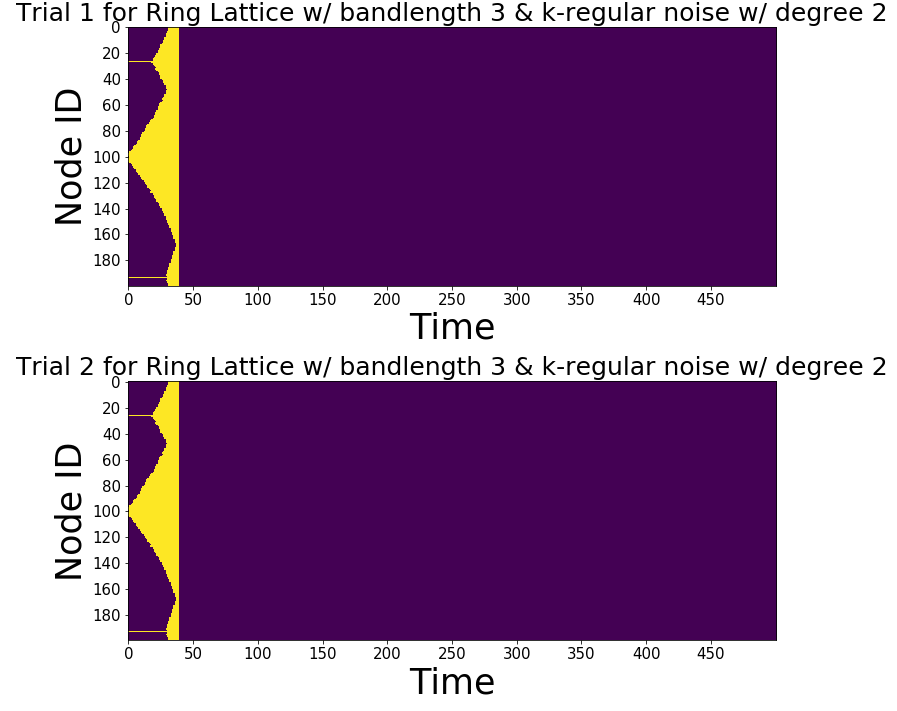
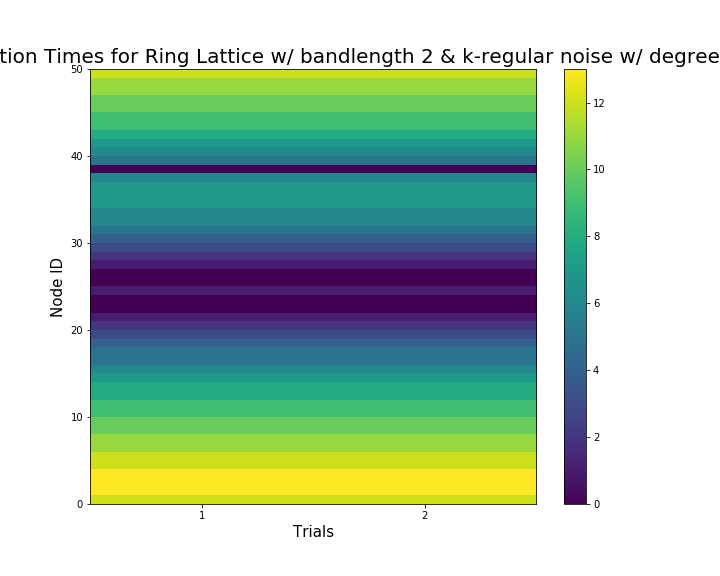
Look at the first activation times¶
Spy activation of the nodes.
ring_latt_k_regular.spy_first_activation(first_excitation_times)
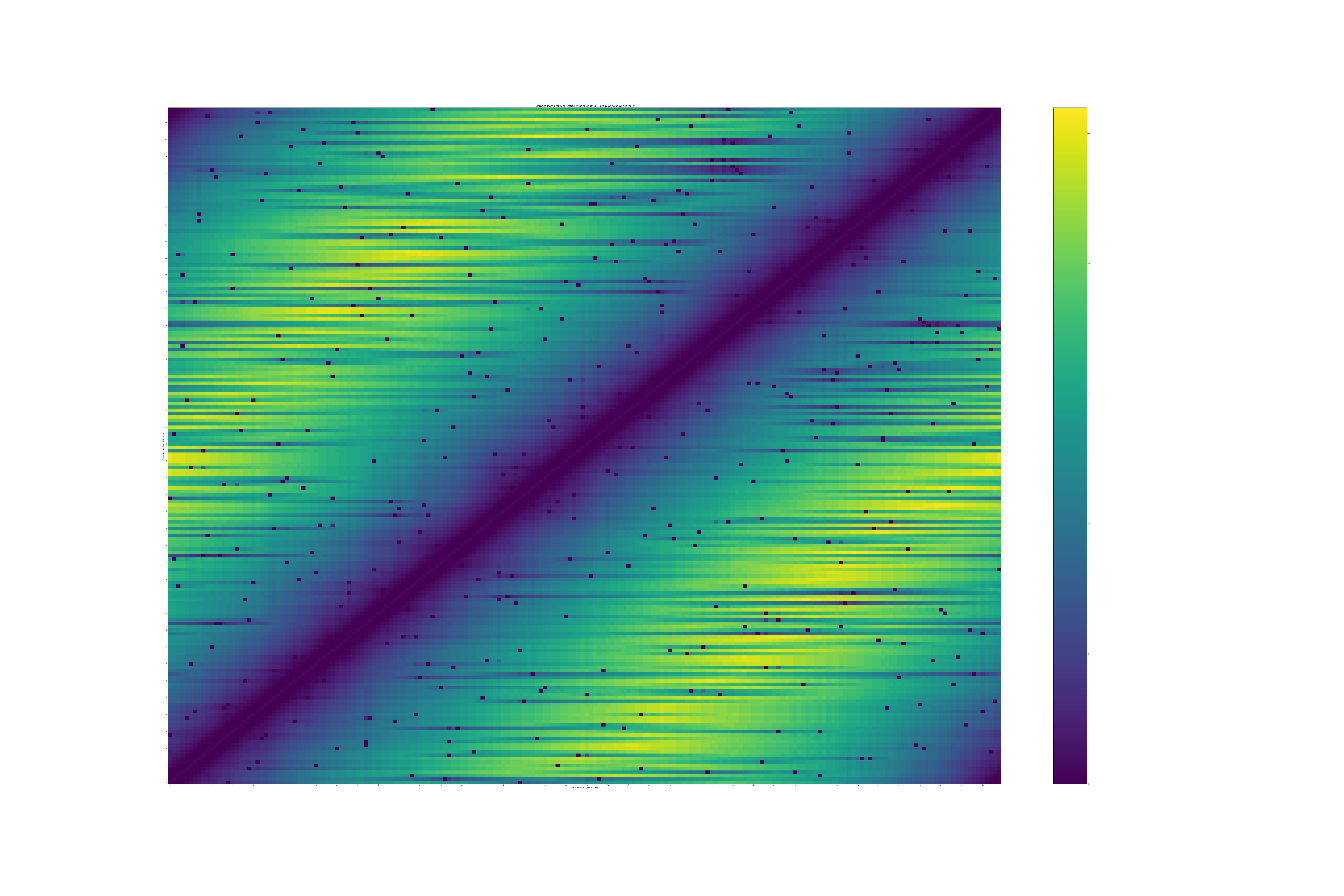
Create Distance Matrix¶
If you don’t need to look at the individual contagions starting from different nodes, you can run the contagion starting from node i and calculating the first time it reaches to node j i.e. create a distance matrix who (i,j) entry is the first time the node j activated on a contagion starting from i.
D, Q = ring_latt_k_regular.make_distance_matrix(T, C, threshold, Trials, refractory_period, spy_distance = True)
Contagion Size¶
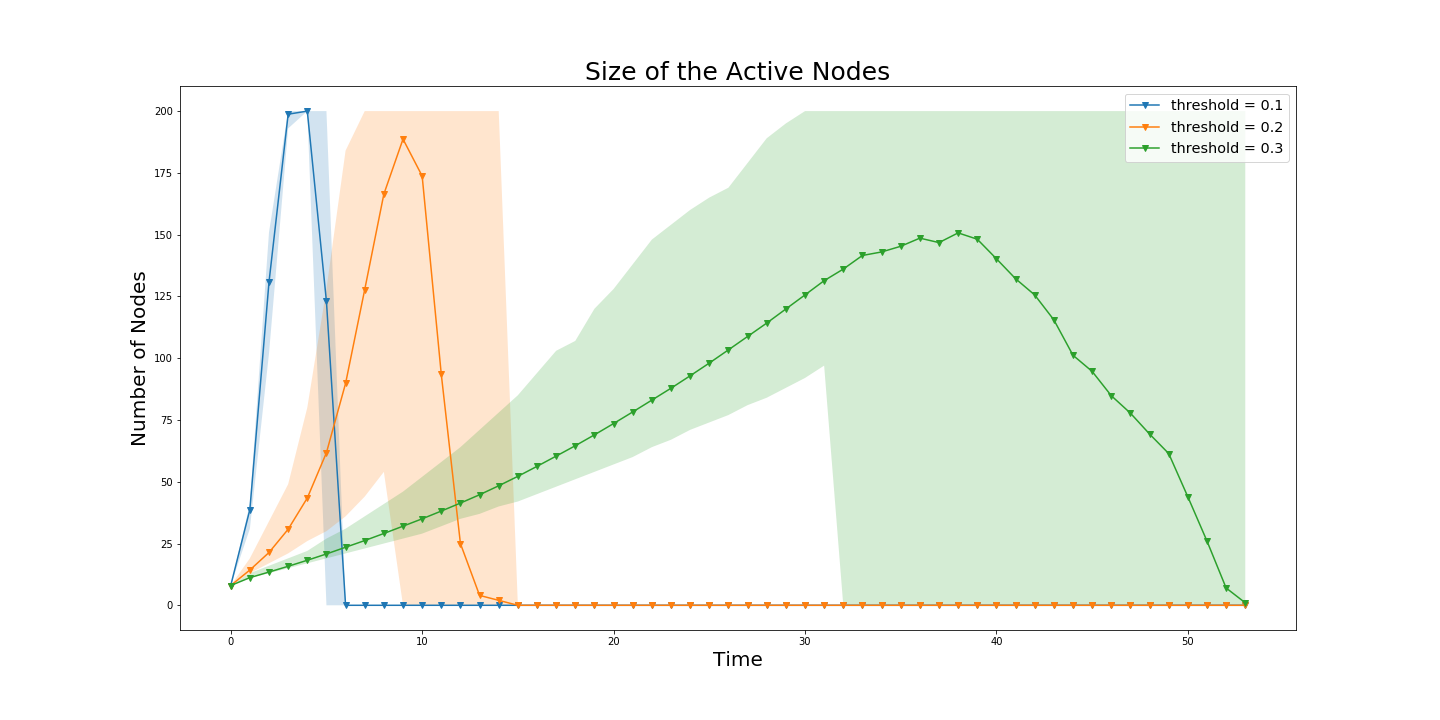
It’s important to look at the bifurcations in the system. In order to do so, one might need to look at the size of the contagion for example different thresholds.
labels = ['threshold = 0.1', 'threshold = 0.2', 'threshold = 0.3']
Q = [Q1,Q2,Q3] ## Qi is the second output of the ``make_distance_matrix``
ring_latt_k_regular.display_comm_sizes(Q,labels)
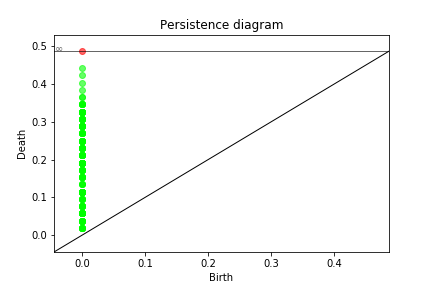
Persistence Diagrams¶
Once we created the distance matrices, we can look at the topological features across different contagions and different topologies.
pers = ring_latt_k_regular.compute_persistence(D, spy = True)
delta = ring_latt_k_regular.one_d_Delta(pers)